A recent study in the field of fluid analysis reveals a diverse set of strategies utilized by alpha and beta variants of the severe acute respiratory syndrome coronavirus 2 (SARS‑CoV‑2) in order to evade antibodies generated during an earlier infection with this virus. The paper is currently available on the bioRxiv* preprint server while it undergoes a peer-review process.
The emergence of SARS-CoV-2 variants of concern – primarily UK B.1.1.7 (known as the alpha variant) and South African B.1.351 (known as the beta variant) – have raised grave concerns that mutations in their spike glycoprotein sequences will improve transmission and generate resistance to multiple neutralizing antibodies.
Hence, understanding the exact factors that enable antibody escape of SARS-CoV-2 and its variants, known to cause coronavirus disease 2019 (COVID-19), is pivotal for the development of effective drugs and vaccines that will bestow broad protection.
Sequencing results have shown that alpha and beta SARS-CoV-2 variants share an N501Y mutation which plays a pivotal role in the interaction between the viral spike glycoprotein and the angiotensin-converting enzyme 2 (ACE2) receptor in human cells.
In contrast to the alpha variant, the beta variant harbors additional E484K and K417N mutations at the key part of spike glycoprotein known as the receptor-binding domain (RBD), raising additional concerns related to viral antibody escape.
By using microfluidic diffusional sizing, a research group led by Dr. Sebastian Fiedler from the United Kingdom resolved the dissociation constant for the interaction between RBDs of SARS-CoV-2 in its original version (also known as wild type virus) and its alpha and beta variants with ACE2 host cell receptor.
A new approach to antibody affinity assessment
In this study, the researchers have used wild-type, alpha and beta RBDs to quantify the impact of various mutations in that exact region (more specifically, mutations such as N501Y, E484K and K417N) on the binding affinity to ACE2 and a monoclonal neutralizing antibody raised against the original Wuhan-Hu-1 SARS-CoV-2.
Moreover, this research group has evaluated the same wild-type and variant RBDs in the process known as microfluidic antibody affinity profiling. This was done to appraise the affinity and concentrations of anti-RBD antibodies in convalescent sera from individuals infected with the original SARS-CoV-2 variant.
Such an approach allowed them to discern changes in affinity across the entire antibody population by appraising the loss of binding in a subset of antibodies. In addition, they have pursued a correlation analysis of these affinities and concentrations with the propensity to disrupt complexes between the receptor and viral spike glycoprotein.
Diverse ways to evade the immune response
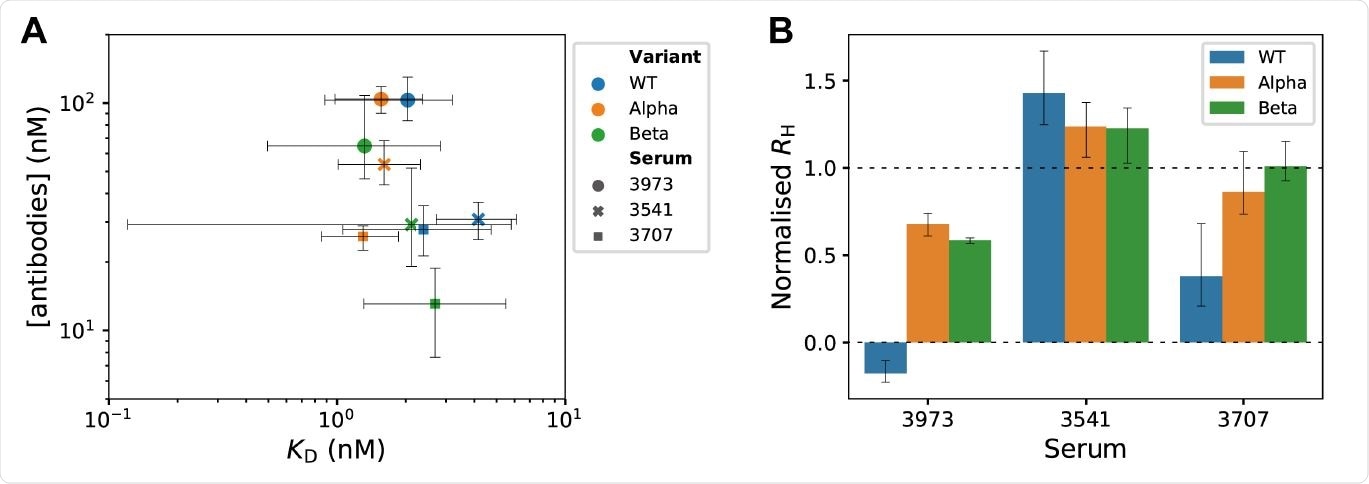
The results show that anti-RBD wild-type antibodies are rather effective against wild-type RBD; however, the effectiveness is much lower for both alpha and beta RBD variants, as they follow different approaches to evade the immune response.
More specifically, for RBD-alpha, antibody escape was primarily driven by an increased binding affinity to the ACE2 receptor. In contrast, RBD-beta seems to halt antibody binding by modifying key epitopes on the protein surface.
As a result, both variants show increased effectiveness in binding to ACE2 on the surface of host cells, increasing, in turn, the prospect of successful cell entry. But, strikingly, the antibody concentration binding to RBD-beta was actually half when compared to RBD-alpha and wild-type RBD.
“Our data, therefore, suggest that one factor contributing to the higher transmissibility and antibody evasion of SARS-CoV-2 alpha and beta is a larger fraction of viruses that can form a complex with ACE2”, say study authors. “However, the two variants employ different mechanisms to achieve this goal,” they add.
Improving our treatment armamentarium
This study has shown that SARS-CoV-2 alpha RBD can bind to ACE2 with greater affinity, which means the displacement from the receptor by neutralizing antibodies is much more cumbersome; however, RBD-beta is less accessible to antibodies due to epitope changes.
In any case, such novel quantitative insights on the complex molecular mechanisms of antibody escape used by SARS-CoV-2 variants of concern are indispensable in understanding how new viral variants actually evolve.
Accordingly, this opens the door for the development of broad therapeutics and vaccine candidates which will be effective against a wide array of current viral variants, and that may be a promising approach against any future viral mutants.
*Important Notice
bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
- Fiedler, S. et al. (2021). Mutations in two SARS-CoV-2 variants of concern reflect two distinct strategies of antibody escape. bioRxiv. https://doi.org/10.1101/2021.07.23.453327, https://www.biorxiv.org/content/10.1101/2021.07.23.453327v1
Posted in: Medical Research News | Disease/Infection News
Tags: ACE2, Angiotensin, Angiotensin-Converting Enzyme 2, Antibodies, Antibody, Assay, binding affinity, Cell, Coronavirus, Coronavirus Disease COVID-19, Drugs, Enzyme, Glycoprotein, Immune Response, Mutation, Protein, Receptor, Research, Respiratory, SARS, SARS-CoV-2, Severe Acute Respiratory, Severe Acute Respiratory Syndrome, Syndrome, Therapeutics, Vaccine, Virus

Written by
Dr. Tomislav Meštrović
Dr. Tomislav Meštrović is a medical doctor (MD) with a Ph.D. in biomedical and health sciences, specialist in the field of clinical microbiology, and an Assistant Professor at Croatia's youngest university – University North. In addition to his interest in clinical, research and lecturing activities, his immense passion for medical writing and scientific communication goes back to his student days. He enjoys contributing back to the community. In his spare time, Tomislav is a movie buff and an avid traveler.
Source: Read Full Article
